Chapter 8 Next steps
Objectives
Introduce the notion of data containers
Give an overview of the
SummarizedExperiment, extensively used in omics analyses
Data in bioinformatics is often complex. To deal with this, developers define specialised data containers (termed classes) that match the properties of the data they need to handle.
This aspect is central to the Bioconductor10 The Bioconductor was initiated by Robert Gentleman, one of the two creators of the R language. Bioconductor provides tools dedicated to omics data analysis. Bioconductor uses the R statistical programming language, and is open source and open development. project which uses the same core data infrastructure across packages. This certainly contributed to Bioconductor’s success. Bioconductor package developers are advised to make use of existing infrastructure to provide coherence, interoperability and stability to the project as a whole.
To illustrate such an omics data container, we’ll present the SummarizedExperiment
class.
8.1 SummarizedExperiment
The figure below represents the anatomy of SummarizedExperiment.
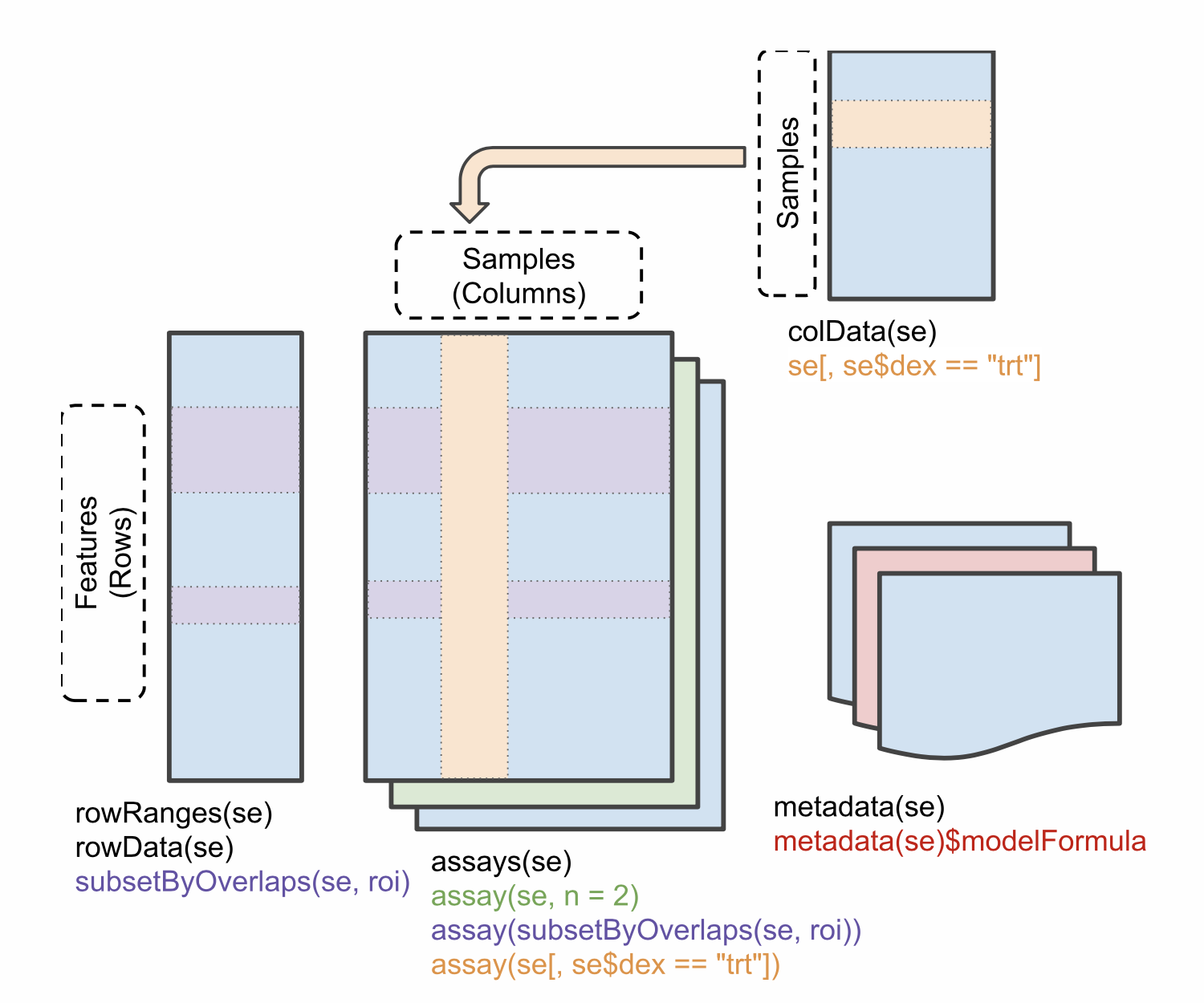
Objects of the class SummarizedExperiment contain :
One (or more) assay(s) containing the quantitative omics data (expression data), stored as a matrix-like object. Features (genes, transcripts, proteins, …) are defined along the rows and samples along the columns.
A sample metadata slot containing sample co-variates, stored as a data frame. Rows from this table represent samples (rows match exactly the columns of the expression data).
A feature metadata slot containing feature co-variates, stored as data frame. The rows of this dataframe’s match exactly the rows of the expression data.
The coordinated nature of the SummarizedExperiment guarantees that during data manipulation, the dimensions of the different slots will always match (i.e the columns in the expression data and then rows in the sample metadata, as well as the rows in the expression data and feature metadata) during data manipulation. For example, if we had to exclude one sample from the assay, it would be automatically removed from the sample metadata in the same operation.
The metadata slots can grow additional co-variates (columns) without affecting the other structures.
8.1.1 Creating a SummarizedExperiment
Remember the rna dataset that we have used previously.
From this table we have already created 3 different tables.
-
An expression matrix: we load the count matrix, specifying that
the first columns contains row/gene names, and convert the
data.frameto amatrix. You can download it by clicking this link.
## GSM2545336 GSM2545337 GSM2545338 GSM2545339 GSM2545340 GSM2545341
## Asl 1170 361 400 586 626 988
## Apod 36194 10347 9173 10620 13021 29594
## Cyp2d22 4060 1616 1603 1901 2171 3349
## Klk6 287 629 641 578 448 195
## GSM2545342 GSM2545343 GSM2545344 GSM2545345 GSM2545346 GSM2545347
## Asl 836 535 586 597 938 1035
## Apod 24959 13668 13230 15868 27769 34301
## Cyp2d22 3122 2008 2254 2277 2985 3452
## Klk6 186 1101 537 567 327 233
## GSM2545348 GSM2545349 GSM2545350 GSM2545351 GSM2545352 GSM2545353
## Asl 494 481 666 937 803 541
## Apod 11258 11812 15816 29242 20415 13682
## Cyp2d22 1883 2014 2417 3678 2920 2216
## Klk6 742 881 828 250 798 710
## GSM2545354 GSM2545362 GSM2545363 GSM2545380
## Asl 473 748 576 1192
## Apod 11088 15916 11166 38148
## Cyp2d22 1821 2842 2011 4019
## Klk6 894 501 598 259
## [ reached 'max' / getOption("max.print") -- omitted 1 row ]## [1] 1474 22- A table describing the samples, available at this link.
## sample organism age sex infection strain time tissue mouse
## 1 GSM2545336 Mus musculus 8 Female InfluenzaA C57BL/6 8 Cerebellum 14
## 2 GSM2545337 Mus musculus 8 Female NonInfected C57BL/6 0 Cerebellum 9
## 3 GSM2545338 Mus musculus 8 Female NonInfected C57BL/6 0 Cerebellum 10
## 4 GSM2545339 Mus musculus 8 Female InfluenzaA C57BL/6 4 Cerebellum 15
## 5 GSM2545340 Mus musculus 8 Male InfluenzaA C57BL/6 4 Cerebellum 18
## 6 GSM2545341 Mus musculus 8 Male InfluenzaA C57BL/6 8 Cerebellum 6
## 7 GSM2545342 Mus musculus 8 Female InfluenzaA C57BL/6 8 Cerebellum 5
## 8 GSM2545343 Mus musculus 8 Male NonInfected C57BL/6 0 Cerebellum 11
## 9 GSM2545344 Mus musculus 8 Female InfluenzaA C57BL/6 4 Cerebellum 22
## 10 GSM2545345 Mus musculus 8 Male InfluenzaA C57BL/6 4 Cerebellum 13
## 11 GSM2545346 Mus musculus 8 Male InfluenzaA C57BL/6 8 Cerebellum 23
## [ reached 'max' / getOption("max.print") -- omitted 11 rows ]## [1] 22 9- A table describing the genes, available at this link.
## gene ENTREZID
## 1 Asl 109900
## 2 Apod 11815
## 3 Cyp2d22 56448
## 4 Klk6 19144
## 5 Fcrls 80891
## 6 Slc2a4 20528
## 7 Exd2 97827
## 8 Gjc2 118454
## 9 Plp1 18823
## 10 Gnb4 14696
## product
## 1 argininosuccinate lyase, transcript variant X1
## 2 apolipoprotein D, transcript variant 3
## 3 cytochrome P450, family 2, subfamily d, polypeptide 22, transcript variant 2
## 4 kallikrein related-peptidase 6, transcript variant 2
## 5 Fc receptor-like S, scavenger receptor, transcript variant X1
## 6 solute carrier family 2 (facilitated glucose transporter), member 4
## 7 exonuclease 3'-5' domain containing 2
## 8 gap junction protein, gamma 2, transcript variant 1
## 9 proteolipid protein (myelin) 1, transcript variant 1
## 10 guanine nucleotide binding protein (G protein), beta 4, transcript variant X2
## ensembl_gene_id
## 1 ENSMUSG00000025533
## 2 ENSMUSG00000022548
## 3 ENSMUSG00000061740
## 4 ENSMUSG00000050063
## 5 ENSMUSG00000015852
## 6 ENSMUSG00000018566
## 7 ENSMUSG00000032705
## 8 ENSMUSG00000043448
## 9 ENSMUSG00000031425
## 10 ENSMUSG00000027669## [1] 1474 9We will create a SummarizedExperiment from these tables:
The count matrix that will be used as the
assayThe table describing the samples will be used as the sample metadata slot
The table describing the genes will be used as the features metadata slot
To do this we can put the different parts together using the
SummarizedExperiment constructor:
First, we make sure that the samples are in the same order in the count matrix and the sample annotation, and the same for the genes in the count matrix and the gene annotation.
stopifnot(rownames(count_matrix) == gene_metadata$gene)
stopifnot(colnames(count_matrix) == sample_metadata$sample)se <- SummarizedExperiment(assays = count_matrix,
colData = sample_metadata,
rowData = gene_metadata)
se## class: SummarizedExperiment
## dim: 1474 22
## metadata(0):
## assays(1): ''
## rownames(1474): Asl Apod ... Lmx1a Pbx1
## rowData names(9): gene ENTREZID ... phenotype_description
## hsapiens_homolog_associated_gene_name
## colnames(22): GSM2545336 GSM2545337 ... GSM2545363 GSM2545380
## colData names(9): sample organism ... tissue mouseUsing this data structure, we can access the expression matrix with
the assay function:
## GSM2545336 GSM2545337 GSM2545338 GSM2545339 GSM2545340 GSM2545341
## Asl 1170 361 400 586 626 988
## Apod 36194 10347 9173 10620 13021 29594
## Cyp2d22 4060 1616 1603 1901 2171 3349
## Klk6 287 629 641 578 448 195
## GSM2545342 GSM2545343 GSM2545344 GSM2545345 GSM2545346 GSM2545347
## Asl 836 535 586 597 938 1035
## Apod 24959 13668 13230 15868 27769 34301
## Cyp2d22 3122 2008 2254 2277 2985 3452
## Klk6 186 1101 537 567 327 233
## GSM2545348 GSM2545349 GSM2545350 GSM2545351 GSM2545352 GSM2545353
## Asl 494 481 666 937 803 541
## Apod 11258 11812 15816 29242 20415 13682
## Cyp2d22 1883 2014 2417 3678 2920 2216
## Klk6 742 881 828 250 798 710
## GSM2545354 GSM2545362 GSM2545363 GSM2545380
## Asl 473 748 576 1192
## Apod 11088 15916 11166 38148
## Cyp2d22 1821 2842 2011 4019
## Klk6 894 501 598 259
## [ reached 'max' / getOption("max.print") -- omitted 2 rows ]## [1] 1474 22We can access the sample metadata using the colData function:
## DataFrame with 22 rows and 9 columns
## sample organism age sex infection
## <character> <character> <integer> <character> <character>
## GSM2545336 GSM2545336 Mus musculus 8 Female InfluenzaA
## GSM2545337 GSM2545337 Mus musculus 8 Female NonInfected
## GSM2545338 GSM2545338 Mus musculus 8 Female NonInfected
## GSM2545339 GSM2545339 Mus musculus 8 Female InfluenzaA
## GSM2545340 GSM2545340 Mus musculus 8 Male InfluenzaA
## ... ... ... ... ... ...
## GSM2545353 GSM2545353 Mus musculus 8 Female NonInfected
## GSM2545354 GSM2545354 Mus musculus 8 Male NonInfected
## GSM2545362 GSM2545362 Mus musculus 8 Female InfluenzaA
## GSM2545363 GSM2545363 Mus musculus 8 Male InfluenzaA
## strain time tissue mouse
## <character> <integer> <character> <integer>
## GSM2545336 C57BL/6 8 Cerebellum 14
## GSM2545337 C57BL/6 0 Cerebellum 9
## GSM2545338 C57BL/6 0 Cerebellum 10
## GSM2545339 C57BL/6 4 Cerebellum 15
## GSM2545340 C57BL/6 4 Cerebellum 18
## ... ... ... ... ...
## GSM2545353 C57BL/6 0 Cerebellum 4
## GSM2545354 C57BL/6 0 Cerebellum 2
## GSM2545362 C57BL/6 4 Cerebellum 20
## GSM2545363 C57BL/6 4 Cerebellum 12
## [ reached 'max' / getOption("max.print") -- omitted 1 row ]## [1] 22 9We can also access the feature metadata using the rowData function:
## DataFrame with 6 rows and 9 columns
## gene ENTREZID product ensembl_gene_id
## <character> <integer> <character> <character>
## Asl Asl 109900 argininosuccinate ly.. ENSMUSG00000025533
## Apod Apod 11815 apolipoprotein D, tr.. ENSMUSG00000022548
## Cyp2d22 Cyp2d22 56448 cytochrome P450, fam.. ENSMUSG00000061740
## Klk6 Klk6 19144 kallikrein related-p.. ENSMUSG00000050063
## Fcrls Fcrls 80891 Fc receptor-like S, .. ENSMUSG00000015852
## Slc2a4 Slc2a4 20528 solute carrier famil.. ENSMUSG00000018566
## external_synonym chromosome_name gene_biotype phenotype_description
## <character> <character> <character> <character>
## Asl 2510006M18Rik 5 protein_coding abnormal circulating..
## Apod NA 16 protein_coding abnormal lipid homeo..
## Cyp2d22 2D22 15 protein_coding abnormal skin morpho..
## Klk6 Bssp 7 protein_coding abnormal cytokine le..
## Fcrls 2810439C17Rik 3 protein_coding decreased CD8-positi..
## Slc2a4 Glut-4 11 protein_coding abnormal circulating..
## hsapiens_homolog_associated_gene_name
## <character>
## Asl ASL
## Apod APOD
## Cyp2d22 CYP2D6
## Klk6 KLK6
## Fcrls FCRL2
## Slc2a4 SLC2A4## [1] 1474 98.1.2 Subsetting a SummarizedExperiment
SummarizedExperiment can be subset just like with data frames, with numerics or with characters of logicals.
Below, we create a new instance of class SummarizedExperiment that contains only the 5 first features for the 3 first samples.
## class: SummarizedExperiment
## dim: 5 3
## metadata(0):
## assays(1): ''
## rownames(5): Asl Apod Cyp2d22 Klk6 Fcrls
## rowData names(9): gene ENTREZID ... phenotype_description
## hsapiens_homolog_associated_gene_name
## colnames(3): GSM2545336 GSM2545337 GSM2545338
## colData names(9): sample organism ... tissue mouse## DataFrame with 3 rows and 9 columns
## sample organism age sex infection
## <character> <character> <integer> <character> <character>
## GSM2545336 GSM2545336 Mus musculus 8 Female InfluenzaA
## GSM2545337 GSM2545337 Mus musculus 8 Female NonInfected
## GSM2545338 GSM2545338 Mus musculus 8 Female NonInfected
## strain time tissue mouse
## <character> <integer> <character> <integer>
## GSM2545336 C57BL/6 8 Cerebellum 14
## GSM2545337 C57BL/6 0 Cerebellum 9
## GSM2545338 C57BL/6 0 Cerebellum 10## DataFrame with 5 rows and 9 columns
## gene ENTREZID product ensembl_gene_id
## <character> <integer> <character> <character>
## Asl Asl 109900 argininosuccinate ly.. ENSMUSG00000025533
## Apod Apod 11815 apolipoprotein D, tr.. ENSMUSG00000022548
## Cyp2d22 Cyp2d22 56448 cytochrome P450, fam.. ENSMUSG00000061740
## Klk6 Klk6 19144 kallikrein related-p.. ENSMUSG00000050063
## Fcrls Fcrls 80891 Fc receptor-like S, .. ENSMUSG00000015852
## external_synonym chromosome_name gene_biotype phenotype_description
## <character> <character> <character> <character>
## Asl 2510006M18Rik 5 protein_coding abnormal circulating..
## Apod NA 16 protein_coding abnormal lipid homeo..
## Cyp2d22 2D22 15 protein_coding abnormal skin morpho..
## Klk6 Bssp 7 protein_coding abnormal cytokine le..
## Fcrls 2810439C17Rik 3 protein_coding decreased CD8-positi..
## hsapiens_homolog_associated_gene_name
## <character>
## Asl ASL
## Apod APOD
## Cyp2d22 CYP2D6
## Klk6 KLK6
## Fcrls FCRL2We can also use the colData() function to subset on something from the sample metadata, or the rowData() to subset on something from the feature metadata.
For example, here we keep only miRNAs and the non infected samples:
## class: SummarizedExperiment
## dim: 7 7
## metadata(0):
## assays(1): ''
## rownames(7): Mir1901 Mir378a ... Mir128-1 Mir7682
## rowData names(9): gene ENTREZID ... phenotype_description
## hsapiens_homolog_associated_gene_name
## colnames(7): GSM2545337 GSM2545338 ... GSM2545353 GSM2545354
## colData names(9): sample organism ... tissue mouse## GSM2545337 GSM2545338 GSM2545343 GSM2545348 GSM2545349 GSM2545353
## Mir1901 45 44 74 55 68 33
## Mir378a 11 7 9 4 12 4
## Mir133b 4 6 5 4 6 7
## Mir30c-2 10 6 16 12 8 17
## Mir149 1 2 0 0 0 0
## Mir128-1 4 1 2 2 1 2
## Mir7682 2 0 4 1 3 5
## GSM2545354
## Mir1901 60
## Mir378a 8
## Mir133b 3
## Mir30c-2 15
## Mir149 2
## Mir128-1 1
## Mir7682 5## DataFrame with 7 rows and 9 columns
## sample organism age sex infection
## <character> <character> <integer> <character> <character>
## GSM2545337 GSM2545337 Mus musculus 8 Female NonInfected
## GSM2545338 GSM2545338 Mus musculus 8 Female NonInfected
## GSM2545343 GSM2545343 Mus musculus 8 Male NonInfected
## GSM2545348 GSM2545348 Mus musculus 8 Female NonInfected
## GSM2545349 GSM2545349 Mus musculus 8 Male NonInfected
## GSM2545353 GSM2545353 Mus musculus 8 Female NonInfected
## GSM2545354 GSM2545354 Mus musculus 8 Male NonInfected
## strain time tissue mouse
## <character> <integer> <character> <integer>
## GSM2545337 C57BL/6 0 Cerebellum 9
## GSM2545338 C57BL/6 0 Cerebellum 10
## GSM2545343 C57BL/6 0 Cerebellum 11
## GSM2545348 C57BL/6 0 Cerebellum 8
## GSM2545349 C57BL/6 0 Cerebellum 7
## GSM2545353 C57BL/6 0 Cerebellum 4
## GSM2545354 C57BL/6 0 Cerebellum 2## DataFrame with 7 rows and 9 columns
## gene ENTREZID product ensembl_gene_id
## <character> <integer> <character> <character>
## Mir1901 Mir1901 100316686 microRNA 1901 ENSMUSG00000084565
## Mir378a Mir378a 723889 microRNA 378a ENSMUSG00000105200
## Mir133b Mir133b 723817 microRNA 133b ENSMUSG00000065480
## Mir30c-2 Mir30c-2 723964 microRNA 30c-2 ENSMUSG00000065567
## Mir149 Mir149 387167 microRNA 149 ENSMUSG00000065470
## Mir128-1 Mir128-1 387147 microRNA 128-1 ENSMUSG00000065520
## Mir7682 Mir7682 102466847 microRNA 7682 ENSMUSG00000106406
## external_synonym chromosome_name gene_biotype phenotype_description
## <character> <character> <character> <character>
## Mir1901 Mirn1901 18 miRNA NA
## Mir378a Mirn378 18 miRNA abnormal mitochondri..
## Mir133b mir 133b 1 miRNA no abnormal phenotyp..
## Mir30c-2 mir 30c-2 1 miRNA NA
## Mir149 Mirn149 1 miRNA increased circulatin..
## Mir128-1 Mirn128 1 miRNA no abnormal phenotyp..
## Mir7682 mmu-mir-7682 1 miRNA NA
## hsapiens_homolog_associated_gene_name
## <character>
## Mir1901 NA
## Mir378a MIR378A
## Mir133b MIR133B
## Mir30c-2 MIR30C2
## Mir149 NA
## Mir128-1 MIR128-1
## Mir7682 NAFor the following exercise, you should download the SE.rda object
(that contains the se object), and open the file using the
‘load()’ function.
download.file(url = "https://github.com/UCLouvain-BIOINFO/bioinfo-training-01-intro-r/raw/refs/heads/main/data/se.rds",
destfile = "data/SE.rds")► Question
Extract the gene expression levels of the 3 first genes in samples at time 0 and at time 8.
► Solution
8.1.2.1 Adding variables to metadata
We can also add information to the metadata. Suppose that you want to add the center where the samples were collected…
## DataFrame with 22 rows and 10 columns
## sample organism age sex infection
## <character> <character> <integer> <character> <character>
## GSM2545336 GSM2545336 Mus musculus 8 Female InfluenzaA
## GSM2545337 GSM2545337 Mus musculus 8 Female NonInfected
## GSM2545338 GSM2545338 Mus musculus 8 Female NonInfected
## GSM2545339 GSM2545339 Mus musculus 8 Female InfluenzaA
## GSM2545340 GSM2545340 Mus musculus 8 Male InfluenzaA
## ... ... ... ... ... ...
## GSM2545353 GSM2545353 Mus musculus 8 Female NonInfected
## GSM2545354 GSM2545354 Mus musculus 8 Male NonInfected
## GSM2545362 GSM2545362 Mus musculus 8 Female InfluenzaA
## strain time tissue mouse center
## <character> <integer> <character> <integer> <character>
## GSM2545336 C57BL/6 8 Cerebellum 14 University of Illinois
## GSM2545337 C57BL/6 0 Cerebellum 9 University of Illinois
## GSM2545338 C57BL/6 0 Cerebellum 10 University of Illinois
## GSM2545339 C57BL/6 4 Cerebellum 15 University of Illinois
## GSM2545340 C57BL/6 4 Cerebellum 18 University of Illinois
## ... ... ... ... ... ...
## GSM2545353 C57BL/6 0 Cerebellum 4 University of Illinois
## GSM2545354 C57BL/6 0 Cerebellum 2 University of Illinois
## GSM2545362 C57BL/6 4 Cerebellum 20 University of Illinois
## [ reached 'max' / getOption("max.print") -- omitted 2 rows ]This illustrates that the metadata slots can grow indefinitely without affecting the other structures!
Take-home message
SummarizedExperimentrepresent an efficient way to store and to handle omics data.They are used in many Bioconductor packages.
If you follow next training focused on RNA sequencing analysis, you will learn to
use the Bioconductor DESeq2 package to do some differential expression analyses.
DESeq2’s whole analysis is handled in a SummarizedExperiment.
Page built: 2025-11-06 using R version 4.5.0 (2025-04-11)